|
Some computer programs can calculate the CV values from ELISA results. Larger variance indicates greater inconsistency and error. This is expressed as a percentage of variance to the mean and indicates any inconsistencies and inaccuracies in the results. The coefficient variation (CV) is the ratio of the standard deviation σ to the mean µ: For these samples, the concentration obtained from the standard curve when analyzing the results must be multiplied by the dilution factor. To obtain an accurate result, these samples should be diluted or concentrated before proceeding with the ELISA staining. Samples that have an absorbance value falling out of the range of the standard curve The concentration of the positive control sample should be within the linear section of the standard curve in order to obtain valid and accurate results. We recommend using a sample of known concentration as a positive control. Each point on the graph represents the mean of the three parallel titrations. We recommend including a standard on each ELISA plate to provide a standard curve for each plate used.Ī representative standard curve is shown in the figure below from the human HIF1 alpha SimpleStep ELISA kit ® (ab171577). Draw a best-fit curve through the points in the graph (we suggest that a suitable computer program be used for this). Duplicates should be within 20% of the mean.Ĭreate a standard curve for the target protein by plotting the mean absorbance (y-axis) against the protein concentration (x-axis). Many computer programs are now available to help process ELISA results in this way.Ĭalculate the average absorbance values for each set of duplicate standards and duplicate samples. This will provide enough data for statistical validation of the results. The maximum absorbance value should be higher than 0.8.Always run ELISA samples in duplicate or triplicate.The OD of the highest standard of a competitive kit should be lower than 0.25.R-squared value is greater than 0.95, and as close to 1 as possible.In general, a good standard curve should have the following characteristics: Schematic diagram of a typical standard curve with trendline, equation and R-squared Addition and formatting of the trendlineįigure 4. This is shown in Figure 4 below.įigure 3. Excel will then display the corresponding trendline formula and R-squared value on the graph. In the formatting section, further select “Display Equation on chart” and “Display R-squared value on chart”. The trendline formatting should then be amended to polynomial order 2 (the default trendline is linear) as shown in Figure 3 below. In Excel, this can be achieved by selecting any point in the XY scatter graph, right-clicking and selecting “Add Trendline”. As shown in Figure 2, the X-axis is the Optical Density, and the Y-axis is the Log of the Standard Concentration.Ī polynomial order 2 trendline can be added to the graph. value of the standard (X-axis) can be plotted against the log of the concentration of the standard (Y-axis), as an XY scatter plot, to get a 5-point curve. 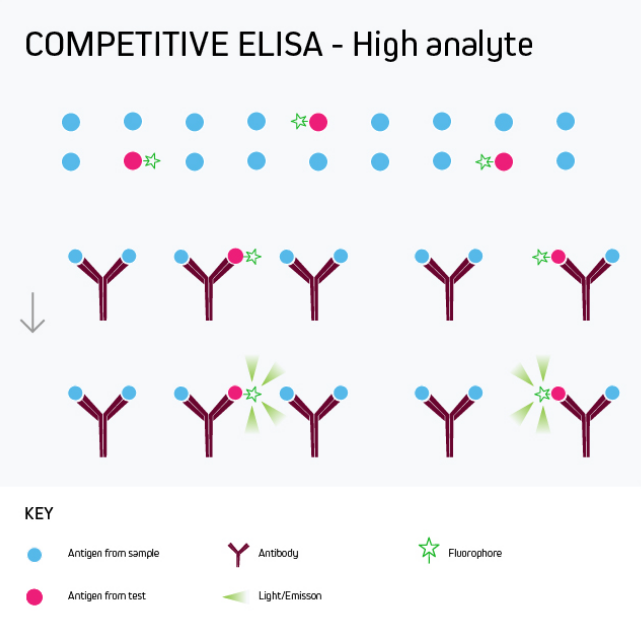
Data of standard concentrations, absorbance values and Log of the standard concentrationįor the competitive inhibition ELISA, there is no need to subtract the absorbance of the negative control. For a competitive inhibition ELISA, the negative control well has the highest OD. As shown in Figure 1, the standard was diluted from 10000 pg/ml to 123.5 pg/ml. In the example below, the standard absorbance values for abx150385, Vitamin B12 ELISA Kit, are shown as a reference. Small molecules such as hormones generally use this method.ĭue to these differences, standard curve fitting is different for sandwich and competitive inhibition ELISAs. 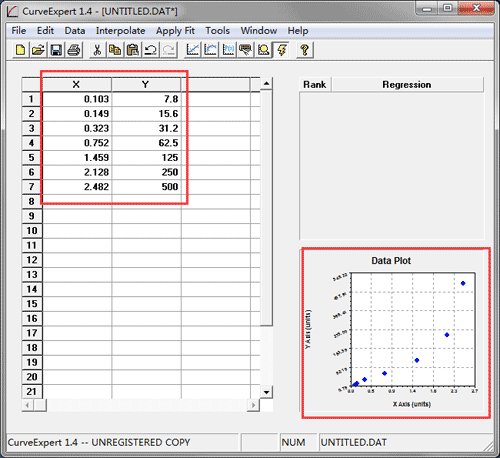
After addition of the substrate solution, the intensity of color developed is inversely proportional to the concentration of target molecules in the samples. This is because the amount of labelled antigen bound to the plate is inversely proportional to the concentration of target molecules in the samples. The higher the sample antigen concentration, the weaker the signal. The concept behind the competitive inhibition ELISA is that free HRP-labelled antigen will compete with the unlabelled antigen in the samples to bind the capture antibodies coated on the plate. In this case, competitive inhibition method can be used to detect the target molecules. Small molecules or haptens do not present more than two binding sites for sandwich detection, and therefore sandwich method is not suitable for detection. 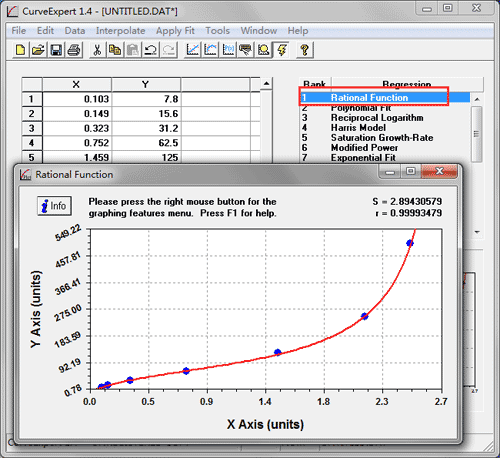
Sandwich ELISA can be applied to the quantitative detection of target antigens or antibodies for antigens which present multiple epitopes. If you are using a non-competitive kit, please view our ELISA Standard Curve Guide. ELISA Competitive Inhibition Standard CurveĮLISA Competitive Inhibition Standard Curve Competitive Inhibition ELISA Standard Curve.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
